Genomic Standards Consortium
The Genomic Standards Consortium (GSC) is an open-membership working body formed in September 2005. The aim of the GSC is making genomic data discoverable. The GSC enables genomic data integration, discovery and comparison through international community-driven standards.
This project is maintained by only1chunts
MIxS Compliance and Implementation
What is metadata?
Metadata is ‘data’ about data. In practical terms, metadata is the information describing a sampling event and subsequent sequencing efforts.
Why use metadata standards?
Utilizing metadata standards to annotate the data describing the sample, sampling environment and sequencing methodology will vastly improve our ability to mine and integrate our sequence data collection for knowledge and application driven research. Collection and reporting of a common, minimal set of metadata across different projects will foster data comparisons and analysis. Combining studies in a standard way will allow for more powerful analyses of data.
Compliance with the MIxS set of standards is very easy – at its minimum it only consists of 11 metadata items, and can be completed very quickly prior to sequence submission to public databases.
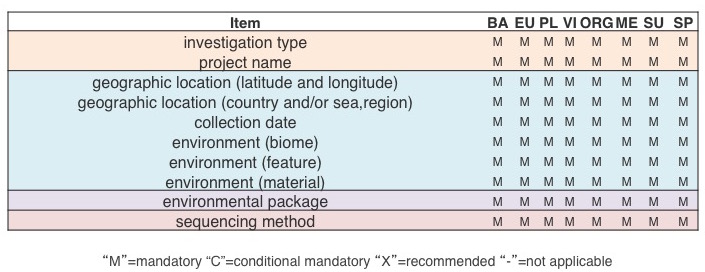
Below are examples of MIxS compliant metadata lists for a genome sequence, a metagenomic sample, and a marker gene survey. They have varying degree of detail, but ultimately what makes them MIxS-compliant are the common items marked in bold red font.
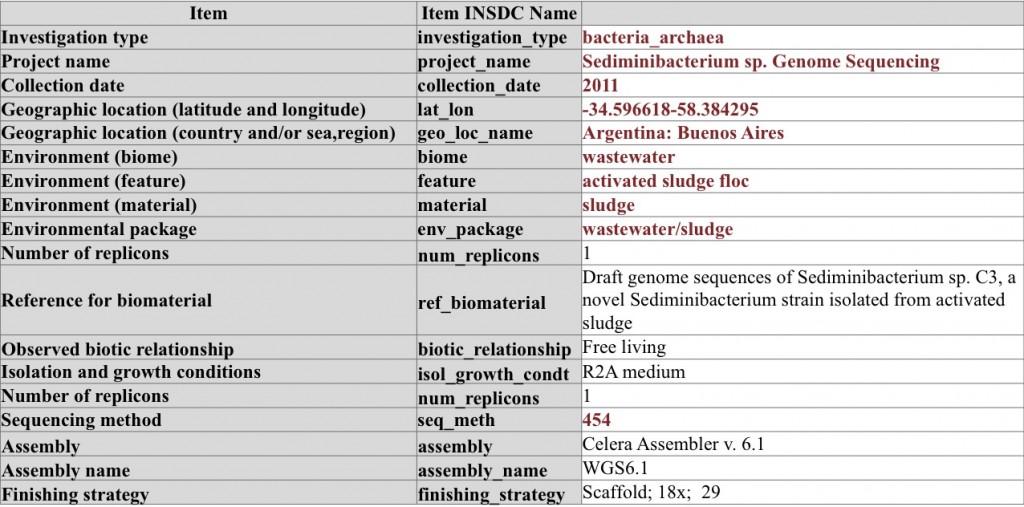
Genome sequencing of Sediminibacterium sp. – note the use of conditional metadata items from the MIGS checklist
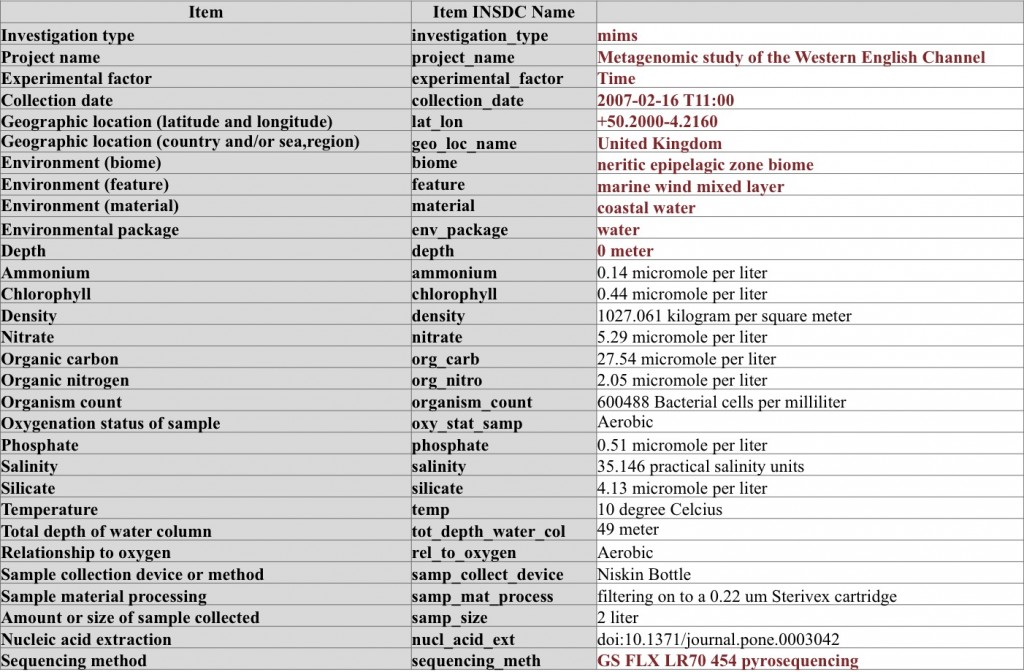
A metagenome (WGS) sequencing sample from sea water – here the sample is extensively characterized by using parameters from environmental package “water”
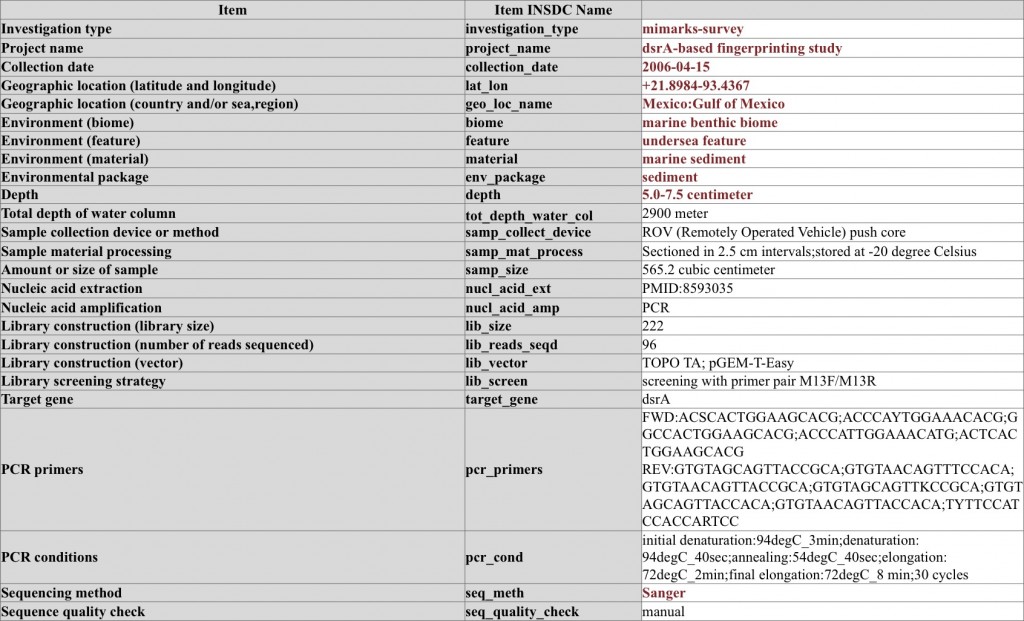
Marker gene survey on dsrA sequences and the accompanying MIMARKS-survey metadata – note the use of MIMARKS checklist conditional mandatory metadata items
Adopters
Despite the relative simplicity of MIxS checklists, it may still not be trivial to prepare the right data in the right format. We compiled a list of databases and tools that help support MIxS to assist submitters further, see the Adopters page.
